
|
|
Projects /
RCTreeDescriptionAdaptive inference describes the problem of repeatedly modifying and performing inference on a model. Since the sequence of models to be used are very similar to one another (i.e., only incremental changes are made at each stage), the results of previous inferential calculations can be used to compute the new results much faster than if performed from scratch.
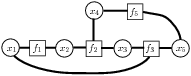 Factor graph 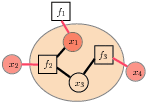 Cluster Factor graphs (left) are used to represent structure in functions,
often probability distributions. Each variable is represented by a variable node (circle), and the overall function is represented as a product of smaller "factors" (squares), whose neighbors are the variables involved in that factor. The notion of a cluster of variables (right) is useful; for a cluster (lightly shaded), its "boundary variables" are the variables whose edges cross the cluster boundary (red).
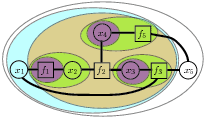 Hierarchical clustering We use a tree contraction process to define a hierarchical clustering of the nodes in the factor graph (left). This clustering then implies a (partial) elimination ordering on the variables in the graph. At each stage of the hierarchy, we compute "cluster functions" which act as sufficient statistics for the nodes below. For marginalization, these functions represent the marginalization of all factors contained in the cluster over all variables except the cluster boundary variables.
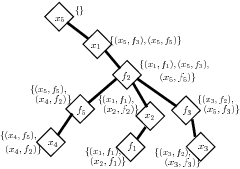 RC-Tree structure Another representation of this hierarchical clustering is an "RC-tree" (short for Rake-and-Compress Tree). Here, each node v of the tree represents a cluster, with children of v representing the (maximal) sub-clusters within v's cluster. Given the cluster functions computed at each node, a query can be computed by passing information down from the root; e.g., the marginal for x2 can be computed using only functions which are direct children of nodes in the path from x2 to the root.
CodeWe have several implementations in Matlab.
|